In offering in vitro drug discovery services to our clients, one of the most common questions we receive is when to use 3D cell culture models and why are they an improvement over traditional 2D cell culture models. It seems obvious that 3D cell culture models would more accurately mimic the in vivo micro environment compared to 2D models, but what are the implications of such a higher degree of in vivo relevance?
In our most recent app note, we performed a side-by-side comparison of the effects of several chemotherapeutic agents on breast cancer cells grown in either 3D spheroid models or via traditional 2D culture methods. First, we found that the Wood breast cancer cell line (derived from an ER +, PR weak+, grade 1, stage T2N0M0 invasive ductal and lobular carcinoma) only expresses ER in 3D, but not 2D culture. Then, we showed that, when dosed with several therapeutics (Paclitaxel, Cisplatin, Carboplatin, Fulvestrant, Tamoxifen, Lapatinib) cell culture models exhibited different sensitivity to drug treatment in 2D versus 3D, particularly in response to Fulvestrant, an ER targeting drug. Therefore, in addition to more accurately representing the physical parameters (e.g. diffusion, gradients) of the in vivo micro environment, 3D cell culture models can also offer a more accurate representation of in vivo gene expression. This improved in vivo relevancy ultimately will enable better translation to in vivo outcomes, thus reducing the cost to develop new therapeutics through more selective and predictive in vitro assays.
Introduction
Breast cancer is one of the most commonly diagnosed cancers among American women, and while early detection and intervention have led to a reduction in the associated mortality rate over the past few decades[1], the development of targeted, patient-specific therapies promises greater success in treating the progressed disease. For example, estrogen receptor (ER), which is expressed by tumors in a subset of breast cancer patients, can facilitate expression of genes responsible for growth and proliferation of breast cancer cells[2], and thus represents an attractive therapeutic target.
Given the rapid pace and growing expense of therapeutic development, high-content in vitro screening approaches represent an attractive approach for assaying target-specific effects of novel drug candidates. However, since cell-extracellular matrix (ECM) interactions, which may not be well represented in traditional 2D cell culture models, are often crucial to the expression of drug targets[3], the application of 2D cell culture models is not always the most effective means to screening targeted therapeutic compounds. Specifically, in the context of breast cancer, several investigations have reported a loss of ER-expression throughout standard 2D culture[4].
In order to explore the effect of 2D versus 3D culture on both ER expression and the consequential response of each culture model to a variety of ER-targeting and non-targeting therapeutics, we implemented the commercially available Wood cell model (Cellaria Biosciences, which was derived from an ER (+), PR (weak+), grade 1, stage T2N0M0 invasive ductal and lobular carcinoma of the breast in a Caucasian female, age 65-69, with a BMI of 33.98. Spheroids cultured in Thermo Fisher multi-well tissue culture plates for 2D adherent culture or Corning® Ultra-low Attachment Spheroid Micro-plates for 3D spheroid culture were dosed with chemotherapeutic agents, labeled with Thermo Fisher’s LIVE/DEAD™ Fixable Red Dead Cell Stain and with Thermo Fisher antibodies against ER and Ki67 (to determine proliferation), and cleared with Visikol® HISTO-M™ for high-content imaging via the Thermo Fisher CX7 LZR. As a result, cell viability and proliferation could be quickly compared across a range of drug treatments in both formats. Herein, we present compelling findings suggesting a difference in expression of ER in 2D versus 3D spheroid culture and consequently a difference in therapeutic response.
Methods
Wood cell culture
Wood cells were maintained in Basal Renaissance Essential Tumor Medium (RETM) with RETM supplement, 25 µg/mL cholera toxin, 5% fetal bovine serum, 1x antibiotic-antimitotic in a humidified, 37°C, 5% CO2 incubator and passaged via light trypsinization with 1X TrypLE and 1 mM EDTA upon reaching 80% confluence. Trypsinized cells were resuspended in complete medium and 1 x 103 cells were plated in each of 384 wells of a Corning Round Bottom Ultra Low Attachment Spheroid Microplate or 2 x 103 cells were plated in each of 96 wells of a Thermo Fisher optically clear, tissue culture treated plate. Spheroids and 2D cell cultures were maintained under standard culture conditions for 2 days, half of the media volume was exchanged to dose with drugs listed in Table 1 in complete media, and cultures were maintained for an additional 2 days prior to labeling.
| Compound | Assayed concentrations | |||||
|---|---|---|---|---|---|---|
| Paclitaxel | 0.01 | 0.1 | 1 | 10 | 100 | nM |
| Cisplatin | 0.1 | 1 | 10 | 100 | 500 | µM |
| Carboplatin | 1 | 10 | 100 | 1000 | 10000 | µM |
| Fulvestrant | 0.01 | 0.1 | 1 | 10 | 100 | nM |
| Tamoxifen | 0.001 | 0.01 | 0.1 | 1 | 10 | µM |
| Lapatinib | 0.01 | 0.1 | 1 | 10 | 100 | µM |
Staining and fixing Wood cells
Cells (spheroids or 2D cultured cells) were washed twice with 1X D-PBS, and a 1:1000 dilution of Thermo Fisher LIVE/DEAD Fixable Red Dead Cell Stain (reconstituted according to manufacturer’s instructions in 1X D-PBS) plus a 1:400 dilution of Molecular Devices Nucview 488 was added to each well. Following 45 minutes of room temperature incubation with the LIVE/DEAD/Nucview 488 staining solution, spheroids/cells were washed twice with 1X D-PBS, fixed with 10% neutral buffered formalin and permeabilized with 0.2% Triton, each for 30 min at room temperature. Fixed spheroids/cells were then blocked with Visikol HISTO Blocking Buffer then labeled with a 1:200 dilution of Ki67 in Visikol HISTO Antibody Buffer, each for 1 h at room temperature. Primary labeled spheroids/cells were then washed with Visikol HISTO 1X Washing Buffer 3 times and labeled with a 1:200 dilution of AlexaFluor 647 anti-rabbit secondary antibody plus a 1:5000 dilution of DAPI in Visikol HISTO Antibody Buffer for 1 h at room temperature. Select spheroids and 2D cell cultures were labeled separately with either DAPI only, AlexaFluor 488 anti-mouse secondary antibody only, or anti-ERα plus AlexaFluor 488 anti-mouse secondary and DAPI only.
Clearing and high throughput imaging of spheroids and 2D cell cultures
Stained spheroids/cells were washed twice with Visikol HISTO Washing Buffer, once with deionized water, once with 50% methanol in deionized water, and once with 100% methanol. For clearing, as much methanol as possible was removed from each well, and Visikol HISTO-M was added for subsequent imaging on a CX7 LZR High Content Confocal Imager. Z-stacks were collected for each spheroid/cell culture (10 µm steps for spheroids, 1 step for 2D cultures). Image analysis was performed using both custom ImageJ macros and CellProfiler.
Results and Discussion
Wood cells exhibit ER expression in 3D but lack detectable levels of ER expression in traditional 2D cell culture
Detectable levels of ER expression were observed in Wood spheroids (A). However, in 2D culture, long exposure times were required to detect any fluorescence in the channel corresponding to ER (B), and fluorescence intensity of ER labeled Wood cells was similar to that of cells labeled with DAPI only (D) or DAPI + secondary only (C), suggesting that ER is not expressed at detectable levels in 2D culture.
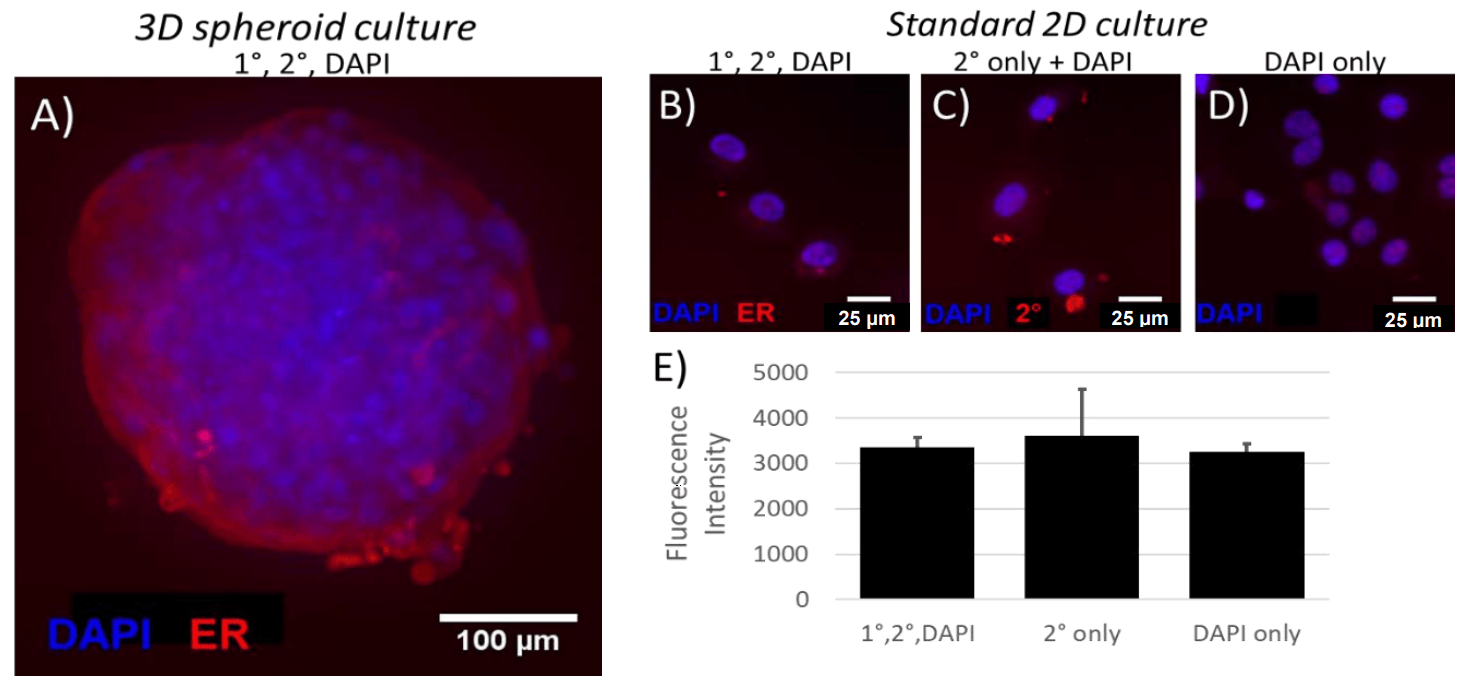
Figure 1: ER expression in 3D spheroids (A) and 2D cultures (B-E). (E) Mean fluorescence intensity of cells from each of 3 labeling conditions. Data represent mean ± SEM; one-way ANOVA with Bonferroni post-hoc revealed no significant differences between 1°/2°/DAPI labeled cells and either 2°-only or DAPI-only labeled cells.
Differences in 2D versus 3D drug response implicate ER expression in spheroid culture in inducing sensitivity to ER-targeting drugs
Treated spheroids (Fig. 2) or 2D cultures (images not shown) were cleared, imaged, and quantitatively analyzed to elucidate the dose response of Wood cells to various drug compounds in each format.
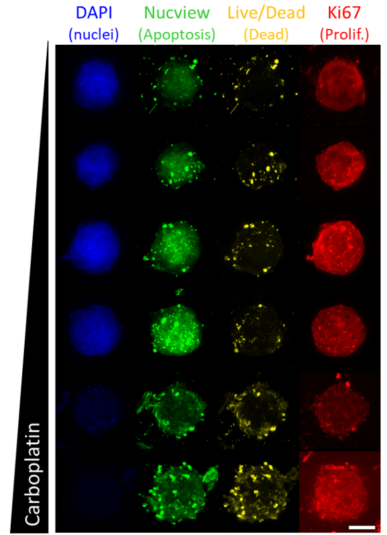
Cells cultured in 3D spheroids exhibit reduced proliferation in proportion to fulvestrant (an ER-targeting drug) dose (Fig. 3), while 2D cultures, which lack detectable ER expression (Fig. 1 B-E) are less sensitive to treatment with fulvestrant (Fig. 3). However, cells cultured in either 2D or 3D exhibit similar sensitivity to carboplatin (Fig. 3), which is not specific to ER, but rather targets DNA synthesis in general[5 ](Fig. 3).
Interestingly, carboplatin treated 2D, but not 3D Wood cells exhibited increased apoptosis and cell death, proportional to carboplatin dose over the range of assayed drug concentrations (Fig. 4A-B). Indeed, carboplatin is known to act primarily by interfering with DNA synthesis[5], implying its relevance in the treatment of breast cancer regardless of hormone receptor status[6]. However, 2D models often exhibit greater sensitivity to general therapeutic treatment, since, in standard 2D culture, drug compounds need not penetrate the dense ECM and multiple cell layers present in spheroid or larger 3D tissues[3]. Therefore, our data suggest the importance of model choice (2D versus 3D) not only in screening targeted effects of therapeutic strategies, but also in assaying general dose-sensitivity, even in the context of non-targeted treatments.
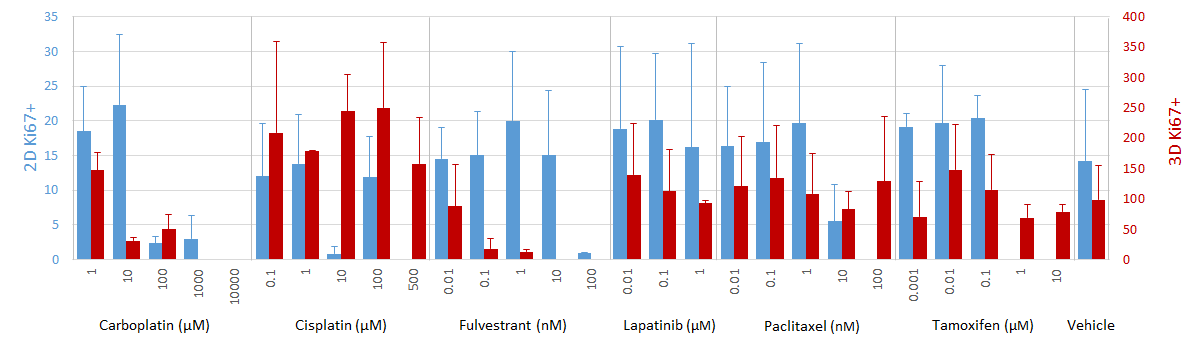
Ki67 response to select drug compounds. Data represent mean ± SD.
Finally, since Tamoxifen is a prodrug, who’s demethylated and hydroxylated metabolites are thought to act as ER antagonists[7], it is expected that the absence of hepatic metabolism in any in vitro models would undermine tamoxifen activity, regardless of 2D or 3D culture format. However, we found that 2D cultures are nevertheless sensitive to tamoxifen treatment (Fig. 4A). Given the lack of detectable ER expression in this culture format (Fig. 1B-E), this suggests that an ER-independent mechanism of action in the context of breast cancer may exist. Indeed, there is evidence in the literature that ER-negative patients respond to tamoxifen treatment[8], underscoring the implications of a potential ER-independent mechanism of action.
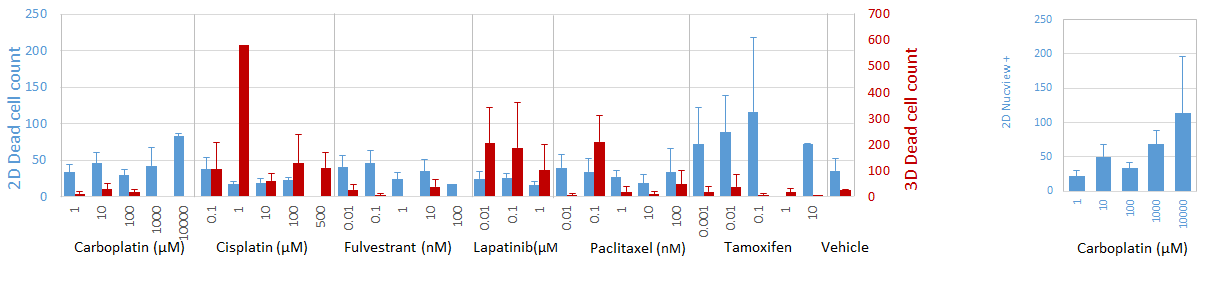
Figure 4: Apoptosis and viability response to select drug compounds. Data represent mean ± SD of dead cells, determined by LIVE/DEAD stain positivity (A) or apoptotic cells, determined by Nucview positivity (B).
All together, these data suggest that not only is model choice crucial to recapitulating targeted effects of chemotherapeutic agents, but comparison of in vitro model formats enables for the generation of mechanistic hypotheses regarding the mechanism of action of targeted and non-targeted therapeutic strategies.
Conclusion
- 3D spheroid and standard 2D cultures of Wood cells exhibit different ER expression and consequently different sensitivity to targeted and non-targeted chemotherapeutic agents, underscoring the importance of model choice (2D versus 3D) in drug screening applications.
- Visikol HISTO-M facilitates clear visualization of the interior of multicellular spheroids, thus enabling the detection and quantification of cell viability and proliferation in the context of 3D breast cancer models.
| Product | Vendor | Cat. No |
|---|---|---|
| 96-well Round Bottom Ultra Low Attachment Spheroid Microplate | Corning | 4515 |
| HepG2 cells | ATCC | HB-8065 |
| Advanced DMEM | Gibco/Thermo Fisher Scientific | 12491015 |
| Fetal Bovine Serum | Gibco/Thermo Fisher Scientific | 10437010 |
| GlutaMAX™ Supplement | Gibco/Thermo Fisher Scientific | 35050061 |
| Antibiotic-Antimycotic (100X) | Gibco/Thermo Fisher Scientific | 15240062 |
| TrypLE Express with Phenol Red | Gibco/Thermo Fisher Scientific | 12605010 |
| EDTA | Fisher Bioreagents | BP248220 |
| DPBS (10X), no calcium, no magnesium | Gibco/Thermo Fisher Scientific | 14200075 |
| Formalin, Buffered, 10% | Fisher Chemical | SF100-20 |
| Triton™ X-100 | Fisher Bioreagents | BP151 |
| DAPI | Sigma | D9542 |
| LIVE/DEAD™ Fixable Green Dead Cell Stain Kit | Molecular Probes/ Thermo Fisher Scientific | L23101 |
| CellInsight™ CX7 LZR High Content Analysis Platform | Thermo Fisher Scientific | CX7B1112LZR |
| Visikol HISTO-M | Visikol | HM-100 |
| Visikol HISTO Antibody Buffer | Visikol | HSK-AB-100 |
| Visikol HISTO Washing Buffer 10X | Visikol | HSK-WB-200 |
References
1. www.breastcancer.org
2. Duffy, M. (2006). Estrogen receptors: role in breast cancer. Critical reviews in clinical laboratory sciences, 43(4), 325-347.
3. Langhans, S., et al. (2018). Three-dimensional in vitro cell culture models in drug discovery and drug repositioning. Frontiers in Pharmacology, 9, 6.
4. Duss, S., et al. (2007). An oestrogen-dependent model of breast cancer created by transformation of normal human mammary epithelial cells. Breast Cancer Research, 9(3), R38.
5. Aggarwal, S. (1993). A histochemical approach to the mechanism of action of cisplatin and its analogues. Journal of Histochemistry & Cytochemistry, 41(7), 1053-1073.
6. Tutt, A, et al. (2018). Carboplatin in BRCA1/2-mutated and triple-negative breast cancer BRCAness subgroups: the TNT Trial. Nature Medicine, 24, 628-637.
7. Dietze, E., et al. (2001). Tamoxifen but not 4-hydroxytamoxifen initiates apoptosis in p53(-) normal human mammary epithelial cells by inducing mitochondrial depolarization. Journal of Biological Chemistry, 276, 5384-5394.
8. Manna, S., et al. (2016). Tamoxifen action on ER-negative breast cancer. Signal Transduction Insights, 5, STI-S29901.

