Highlights
When combined with high-content confocal microscopy and commercially available viability reagents, Visikol® HISTO-M™ allows the interior environment of 3D cell cultures to be imaged, facilitating the detection and quantification of a necrotic core in large HepG2 spheroids. Spheroids larger than 200 µm in diameter present a necrotic core, indicated by higher frequencies of centralized, non-viable cells.
Introduction
Given their low-cost and high-throughput nature, multicellular spheroid models have become a mainstay in high content screening approaches to drug discovery and development in various physiological and pathophysiological contexts. For example, multicellular spheroids have been generated from cancerous cells to screen potential chemotherapeutic agents[1], hepatocytes to screen for drug-related liver toxicity[2], and even isolated primary cells from the human cortex to study drug penetration into the blood brain barrier[3]. Given the crucial nature of cell viability in assaying the cytotoxic effects of therapeutic candidates in these various contexts, a reliable determination of the frequency and distribution of non-viable cells is necessary for the establishment of a baseline in newly developed models and for determining the localization of cytotoxic effects in treated models[1].
Here, we employed the commonly used hepatocyte cell line, HepG2 to generate multi-cellular spheroids of varying sizes using Corning® Ultra-low Attachment Spheroid Micro-plates. Combined with ThermoFisher’s LIVE/DEAD™ Fixable Green Dead Cell Stain and spheroid clearing with Visikol HISTO-M we were able to determine the frequency and localization of non-viable cells.
Methods
HepG2 cell culture
HepG2 cells were maintained in Advanced DMEM, supplemented with 5% fetal bovine serum, 1x GlutaMAX and 1x antibiotic-antimitotic in a humidified, 37°C, 5% CO2 incubator and passaged via light trypsinization with 0.05% trypLE and 1 mM EDTA upon reaching 80% confluence. Trypsinized cells were resuspended in complete medium and 5 x 102 – 16 x 103 cells were plated in each of 96 wells of a Corning Round Bottom Ultra Low Attachment Spheroid Microplate. Spheroids were maintained under standard culture conditions for 4 days, exchanging half of the media volume with fresh complete media after 2 days.
Staining and fixing HepG2 spheroids
Spheroids were washed twice with 1X D-PBS, and a 1:1000 dilution of Thermo LIVE/DEAD Fixable Green Dead Cell Stain (reconstituted according to manufacturer’s instructions in 1X D-PBS) was added to each well. Following 45 minutes of room temperature incubation with the LIVE/DEAD staining solution, spheroids were washed twice with 1X D-PBS, fixed with 10% neutral buffered formalin and permeabilized with 0.2% Triton, each for 30 min at room temperature. Fixed spheroids were then stained with a 1:5000 dilution of DAPI in Visikol HISTO Antibody Buffer for 1 h at room temperature.
Clearing and high throughput imaging of spheroids
Stained spheroids were washed twice with Visikol HISTO washing buffer, once with deionized water, once with 50% methanol in deionized water, and once with 100% methanol. For clearing, as much methanol as possible was removed from the well, and Visikol HISTO-M was added to each well for subsequent imaging on a CX7 LZR High Content Confocal Imager. Z-stacks were collected for each spheroid, using 10 µm steps. Image analysis was performed using both custom ImageJ macros and CellProfiler.
Results and Discussion
Higher cell seeding densities result in larger spheroids
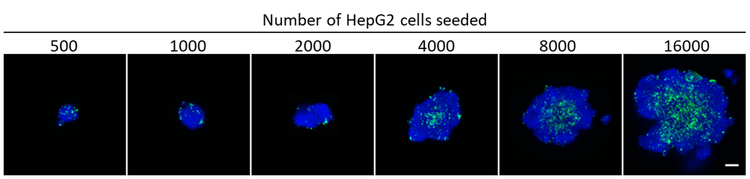
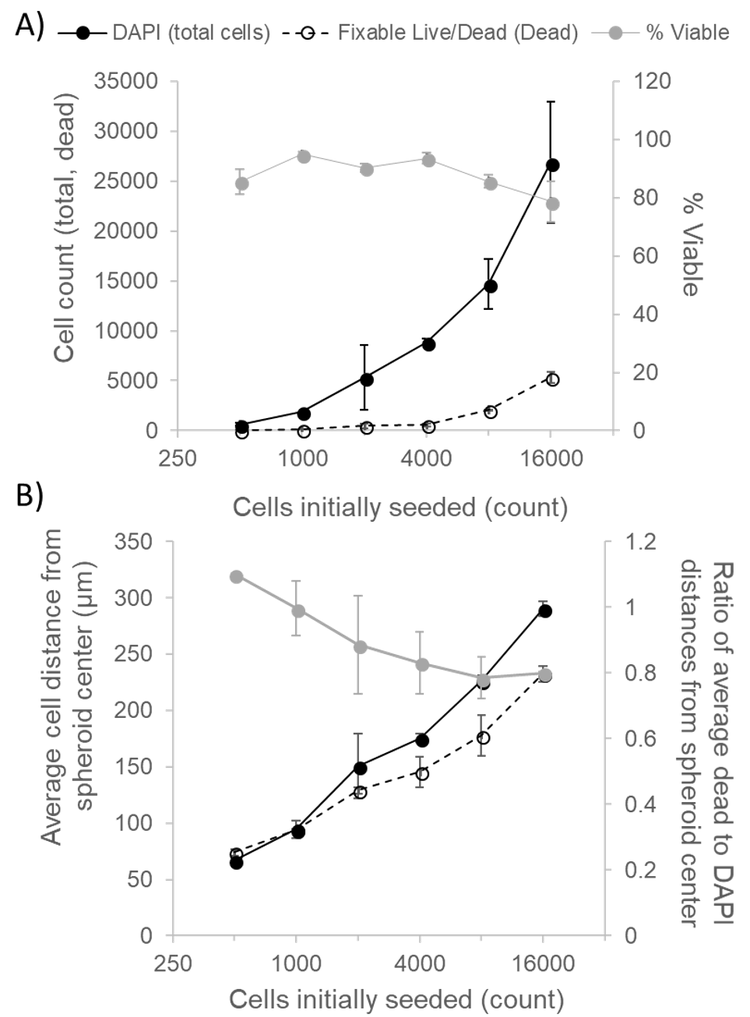
As seen in Figure 1, higher cell seeding densities resulted in larger, more irregularly-shaped spheroids. Quantifying DAPI and Fixable Dead Cell Stain-positive cells using CellProfiler revealed an increase in both the number of total and dead cells with increasing cell seeding densities (Fig. 2A).
Larger spheroids result in the appearance of a necrotic core
While the percentage of viable cells did not precipitously drop with increasing cell seeding densities (Fig. 2A), the localization of non-viable cells qualitatively differed (Fig. 1). Accordingly, we quantified the mean distances from the spheroid center of each DAPI and Fixable Dead Cell Stain-positive cells. As expected, the average distance from the center of both total and dead cells increased with increasing seeding density (Fig. 2B). However, when taking the ratio of the average distance of the dead cells from the spheroid center to the average distance of all cells from the spheroid center, the ratio is inversely proportional to the number of cells seeded, suggesting the development of a necrotic core with spheroid size. The ratio drops below 1 at seeding densities above 1000 cells. Given the diameter of spheroids at this size, this data suggests the development of a necrotic core for HepG2 spheroids larger than 200 µm in diameter.
Spheroid clearing enables clearer visualization and quantitation of a necrotic core
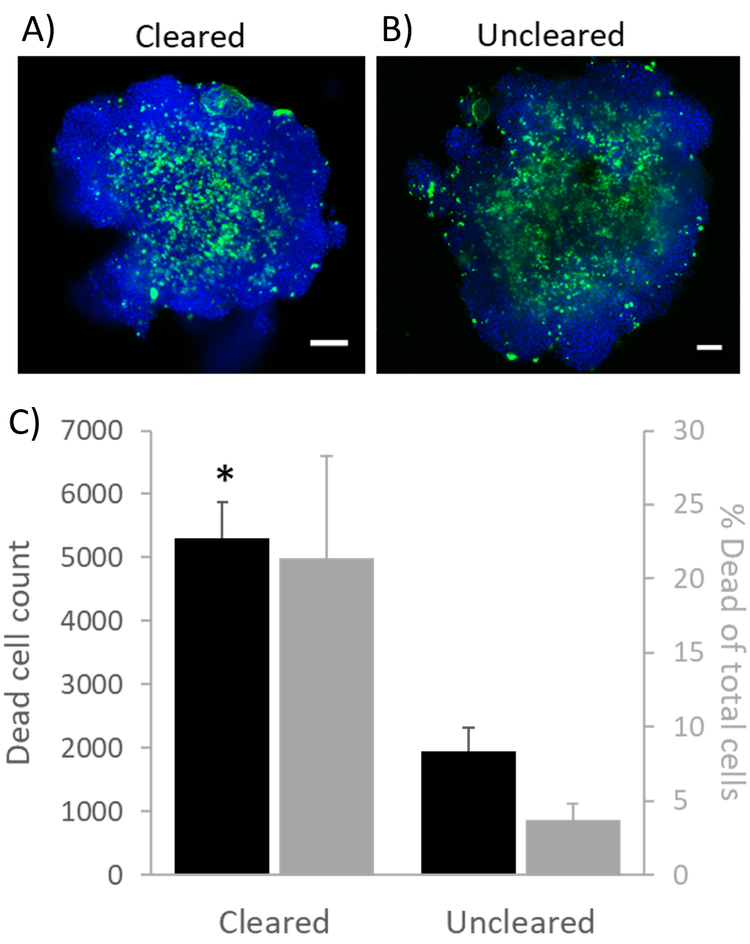
When comparing cleared versus uncleared spheroids generated using the highest seeding density tested (16,000 cells/well), uncleared spheroid centers featured less punctate dead cell staining and an absence of DAPI staining at the core (Fig. 3A-B), thus complicating image analysis. This resulted in misleadingly lower dead cell counts in uncleared relative to cleared spheroids (Fig. 3C).
Conclusions
- Visikol HISTO-M facilitates clear visualization of the spheroid interior, thus enabling the detection, quantification, and localization of non-viable cells.
- Spheroids larger than 200-300 µm in diameter present a necrotic core.
- Detection and quantification of spheroid necrotic cores may prove crucial to the development of larger, more-complex multi-cellular spheroid models for high-throughput drug screening assays.
| Product | Vendor | Cat. No |
|---|---|---|
| 96-well Round Bottom Ultra Low Attachment Spheroid Microplate | Corning | 4515 |
| HepG2 cells | ATCC | HB-8065 |
| Advanced DMEM | Gibco/Thermo Fisher Scientific | 12491015 |
| Fetal Bovine Serum | Gibco/Thermo Fisher Scientific | 10437010 |
| GlutaMAX™ Supplement | Gibco/Thermo Fisher Scientific | 35050061 |
| Antibiotic-Antimycotic (100X) | Gibco/Thermo Fisher Scientific | 15240062 |
| TrypLE Express with Phenol Red | Gibco/Thermo Fisher Scientific | 12605010 |
| EDTA | Fisher Bioreagents | BP248220 |
| DPBS (10X), no calcium, no magnesium | Gibco/Thermo Fisher Scientific | 14200075 |
| Formalin, Buffered, 10% | Fisher Chemical | SF100-20 |
| Triton™ X-100 | Fisher Bioreagents | BP151 |
| DAPI | Sigma | D9542 |
| LIVE/DEAD™ Fixable Green Dead Cell Stain Kit | Molecular Probes/ Thermo Fisher Scientific | L23101 |
| CellInsight™ CX7 LZR High Content Analysis Platform | Thermo Fisher Scientific | CX7B1112LZR |
| Visikol HISTO-M | Visikol | HM-100 |
| Visikol HISTO Antibody Buffer | Visikol | HSK-AB-100 |
| Visikol HISTO Washing Buffer 10X | Visikol | HSK-WB-200 |
Figure 2 (A) Total and dead cell counts and percent viability of HepG2 spheroids, determined from confocal z-stacks. (B) Mean distances of total cells and dead cells, calculated from spheroid center as well as ratio of these distances. Data represent mean values ± SEM for independently imaged spheroids.

Figure 3 Confocal z-slices of (A) cleared and (B) uncleared HepG2 spheroids. (C) Quantification of dead cell frequencies and percentages of dead of total (DAPI labeled) cells. Blue = DAPI; Green = LIVE/DEAD Fixable Green Dead Cell Stain. Scale bar 100 µm. Data represent mean values ± SEM for independently imaged spheroids; * indicates p<0.05 for two-tailed two-sample t-test between cleared and uncleared means.
References
1. Zanoni, M., Piccinini, F., Arienti, C., et al. (2016). 3D Tumor Spheroid Models for In Vitro Therapeutic Screening: A Systematic Approach to Enhance the Biological Relevance of Data Obtained. Scientific Reports, 6(1), 19103.
2. Gunness, P., Mueller, D., Shevchenko, V., et al. (2013) 3D Organotypic Cultures of Human HepaRG Cells: A Tool for In Vitro Toxicity Studies. Toxicolocial Sciences, 133(1), 67-78.
3. Nzou, G., Wicks, R.T., Wicks, E.E., et al. (2018). Human Cortex Spheroid with a Functional Blood Brain Barrier for High-Throughput Neurotoxicity Screening and Disease Modeling. Scientific Reports, 8(1), 7413.

