RT-PCR, the laboratory technique that adds reverse transcription to the already game-changing polymerase chain reaction technology, can be overwhelming to tackle for the first time. Usually employed in gene expression assays, RT-PCR offers researchers the chance to explore the RNA within a sample via reverse transcription into complementary DNA (cDNA), followed by PCR – often quantitative (qPCR) – targeting one or more genes of interest. There are a number of choices to be made in planning these experiments, from RNA extraction protocol to quantitative PCR method. One of the most critical decisions to be made is whether to follow a one-step or two-step procedure.
The distinction between the two is fairly simple. In one-step RT-PCR, both major reactions (reverse transcription and polymerase chain reaction) are carried out in a single tube. For a two-step process, the cDNA products of reverse transcription are transferred to a second tube for subsequent PCR. For multiple qPCR assays, this might actually involve a number of separate reactions, and thus separate vessels with which to work. While a quick glance at these procedures may make the one-step protocol seem like an obvious choice, the decision is actually a lot more nuanced.
One-step RT-PCR
One-step RT-PCR does have the advantage of being just that: one step! Not only is workflow simplified, but keeping everything in a single tube for both major steps reduces the chances for human error to impact your experimental results. Pipetting variance has a smaller effect when you can eliminate some of the pipetting! This can be a very efficient way to move from RNA to expression results for one or even a few genes.
However, one-step RT-PCR has some major limitations. Most importantly, you are limited to the PCR reactions that can be carried out in a single tube. If you plan to target multiple genes with a qPCR method based upon measuring double-stranded DNA, or if you have more targets than can be achieved in a single multiplex reaction, one-step RT-PCR probably isn’t the most efficient way to go about it.
Two-step RT-PCR
Choosing two-step RT-PCR, on the other hand, allows you to immediately move from RNA to the more stable cDNA without even having to know exactly which genes you will be targeting downstream. A full range of targets can be chosen after seeing preliminary qPCR results. You can even hold onto cDNA samples indefinitely, returning to them well into the future to look for expression of genes you may not have initially considered.
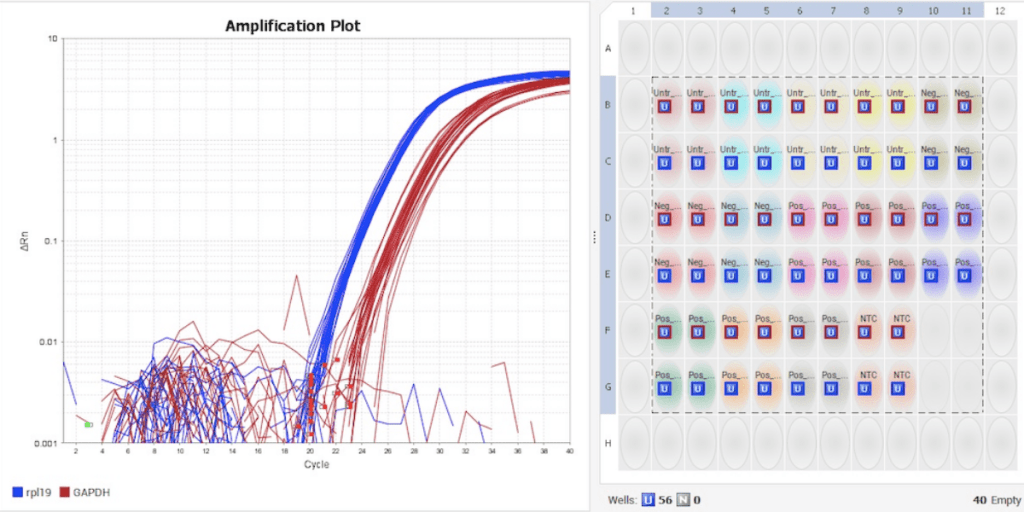
Visikol usually utilizes the two-step method for work examining gene expression. Many of Visikol’s clients choose SYBR Green-based qPCR for their assays and have a number of gene targets in mind. Two-step RT-PCR allows for a quick move from RNA to cDNA, which can then be used as needed in subsequent qPCR reactions. Target genes may be added to the project after analyzing preliminary PCR experiments, providing a degree of flexibility not available with the one-step protocol.
However, Visikol is committed to making experimental design decisions that balance cost, efficiency, and the specific endpoint needs of the client, and so openness to the workflow of both one- and two-step RT-PCR is important. If you have RT-PCR experiments in mind for your drug discovery or disease progression assay, feel free to get in touch with us to discuss your options with a member of our experienced team.
