Visikol leverages its expertise in the field of advanced imaging and image analysis – with special emphasis on 3D – to execute service projects for Clients interested in analyzing organs, tissues, and tissue sections in greater qualitative and quantitative detail. Traditionally, tissues have been characterized through the process of embedding, sectioning, staining and traditional brightfield microscopy. While this is the paradigm for histological examination, it limits analysis to small subregions of tissue, and falls short in the analysis of heterogeneous and complex tissues, such as certain solid tumors or brain tissue. For example, complex three-dimensional features such as vasculature are challenging to evaluate using traditional two-dimensional histology.
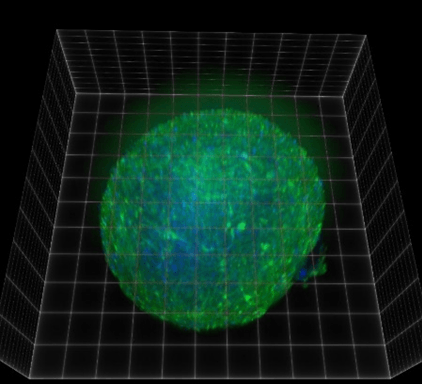

Figure 1. 3D view of iPSC-derived cortical spheroid immunolabeled for GFAP-expressing astrocytes (green) with DAPI nuclear counterstain (blue)
At Visikol, small molecule probes and immunofluorescent labeling are combined with our patented Visikol® HISTO™ tissue clearing approach, and by utilizing confocal microscopy or light sheet imaging, we can image thick tissue specimens in 3D. This allows us to capture the entire population of cells throughout the depth of a tissue instead of being limited to thin two-dimensional sections. The ability to image tissues in 3D enables novel research questions to be answered, which is especially useful in neuroscience, cancer biology, and the study of complex organ systems.
We provide end-to-end 3D tissue imaging services that include tissue processing, labeling, imaging, image analysis and reporting. We were the first company to offer 3D tissue imaging services to Clients and currently provide confocal microscopy and light sheet imaging to our Clients depending on the type of research question and desired endpoints. While we can accommodate whole organ labeling, clearing and imaging, we find that the most optimal approach for most research questions is to cut tissues into approx. 1-2 mm thick sections for processing and imaging and then to conduct imaging with high-content automated confocal microscopy. If needed, the resulting image stacks can be stitched back together digitally. The process we use minimizes labeling and imaging time, which lowers costs for our clients. The resulting image data is delivered to Clients via hard-drive or the cloud – depending on preference. Typically, the 3D data sets that we generate are very large (potentially 100’s of gigabytes) and unless a researcher is equipped with the appropriate software and hardware to handle the load, the analysis can be quite challenging. We use our 3Screen™ image analysis platform, specifically built for large 3D datasets, to analyze this data, and we deliver actionable, easy-to-digest reports.
Our imaging services are offered to Clients in a variety of formats. For example, we can receive fixed tissues for running a more all-encompassing study that involves label optimization along with imaging and image analysis or we can take already generated image sets and run image analysis exclusively using our 3Screen™ software. The latter approach is often convenient for large data sets that have limited and mostly qualitative data associated with them. For each Client, we work closely with them to determine the optimal project design. We have worked on a wide range of projects and are agnostic in our approach — whether it be imaging modality, staining techniques, tissue clearing method or tissue type. We see ourselves as an extension of our Client’s research team and work with our Clients to design a robust and effective approach for their specific application.
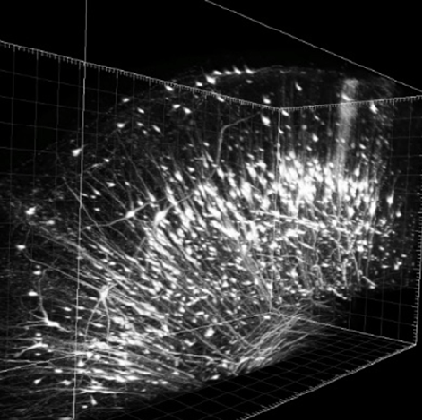
Figure 2. Neurons labeled with rhodamine in the visual cortex of a snowy owl brain.
If you have any questions or would like to learn more, feel free to reach out: info@visikol.com

