Mass Spectrometry (MS) is an important analytical instrument that plays a heavy role in drug discovery and development. Especially in the pharmaceutical industry, it is an essential instrument to properly utilize to identify various compounds in the experiment. MS determines the mass-to-charge ratio of ions in each sample. The techniques involve the ionization of molecules in the sample, followed by the separation and detection of the resulting ions based on their mass-to-charge ratios.
Types of Mass Spectrometry
A few of the various types of MS we use in pharmacology are Tandem Mass Spectrometry (MS/MS) and High-Resolution Mass Spectrometry (HRMS). MS/MS is a two-stage process where there is the initial ionization of the compound called the precursor ion into smaller fragments of ions, and then in the second stage, a selected ion from the first stage is further fragmented in the ion trap or other mass analyzer. The resulting tandem mass spectra provide more detailed information about the structure and allows for the precise quantitative analysis of a compound in each biological sample. This two-stage ionization of a compound allows further analysis of the fragments in both the parent and daughter ion scan, that are unique to the compound.
High resolution mass spectrometry (HRMS) is a type of mass spectrometry that can accurately measure the mass-to-charge ratios of ions with high precision and accuracy. Unlike conventional mass spectrometry, which operates at low resolutions, HRMS is capable of separating and detecting ions that differ in mass by as little as 0.001 atomic mass units (amu). This high level of resolution enables the detection and identification of complex mixtures of molecules with high sensitivity and specificity.
The two modes of measuring compounds via MS are similar, yet very different in application. MS/MS is useful when trying to understand structural information by fragmentations and helps when trying to understand the reactivity of a compound. HRMS is useful when trying to identify a specific compound with precise mass measurements with high resolution and this method is often used when identifying unknown compounds.
Here at Visikol, we use MS to quantify and validate different methods to make a project analysis feasible. When handling unknown compounds, we use HRMS to identify and validate the compounds of interest to be able to, in high accuracy, identify yielded ions that are unique to the compound. After initial analysis, MS/MS quantifications further help in understanding the behavior of the compound.
HUREL® and Mass Spectrometry
One of the ways we effectively use MS is with HUREL, our micro liver system. We use HRMS to identify varying types of metabolites from different HUREL systems. Because all these metabolites have very similar structures but minutely varying mass, HRMS can detect the slightest difference between the metabolites produced. An example is Timolol, a beta-blocker used in many applications, and when it is metabolized with our HUREL systems, it produced 6 metabolites (Figure 1).
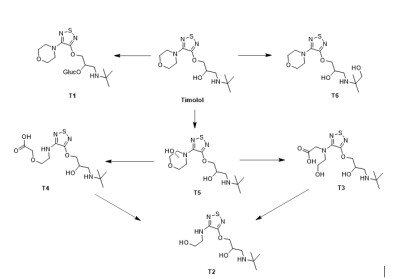
Figure 1: Timolol metabolites generated on HUREL models.
LC-MS/MS
With LC-MS/MS, we were able to understand the clearance of the compound and how the molecules are modified over time in different HUREL hepatic systems. It is important to quantify the samples in a precise and accurate manner because one small difference in a decimal point can shift the identification of the ions in a different manner. As seen in Figure 2, the m/z values of the metabolites are reported in 7 significant figures. Using HRMS was especially useful when differentiating between T3, T4 and T5, T6 as those sets of metabolites had minute mass differences detected in the thousandths. With a low-resolution MS instrument, these sets of compounds would deem identical but with HRMS’s high sensitivity, it was capable of distinguishing between the smallest difference in mass of the metabolites, allowing accurate analysis.
With LC-MS/MS, we were able to understand the clearance of the compound and how the molecules are modified over time in different HUREL hepatic systems. It is important to quantify the samples in a precise and accurate manner because one small difference in a decimal point can shift the identification of the ions in a different manner. As seen in Figure 2, the m/z values of the metabolites are reported in 7 significant figures. Using HRMS was especially useful when differentiating between T3, T4 and T5, T6 as those sets of metabolites had minute mass differences detected in the thousandths. With a low-resolution MS instrument, these sets of compounds would deem identical but with HRMS’s high sensitivity, it was capable of distinguishing between the smallest difference in mass of the metabolites, allowing accurate analysis.
| Code | Metabolite | Observed m/z | HUREL® Human | HUREL® Dog | HUREL® Rat | HUREL® Mouse | HUREL® Minipig |
|---|---|---|---|---|---|---|---|
| Tim | Parent drug | 317.1638 | |||||
| T1 | Tim + Gluc | 493.1956 | Present | Present | Weak | – | – |
| T2 | Tim-C2H2 | 291.1485 | Present | Present | Present | Present | Weak |
| T3 | Tim + 2O | 349.1537 | Present | Present | Present | Present | Present |
| T4 | Tim + 2O | 349.1535 | Present | Present | Present | Present | Present |
| T5 | Tim + O | 333.1587 | Present | – | Weak | Weak | – |
| T6 | Tim + O | 333.1588 | Present | Present | Present | Present | – |
Table 1. Timolol metabolites observed from five HUREL models.
As seen from the chromatogram of the metabolites from the two HUREL models (Rat & Mouse) there are small differences in the peak area of metabolites present in these two systems. Although similar species, the hepatic systems are different in that there is no T1 detected in the HUREL Mouse, but is found in the HUREL Rat. Although T4 is present in both systems, the value obtained from the two systems is different as shown by the peak area.
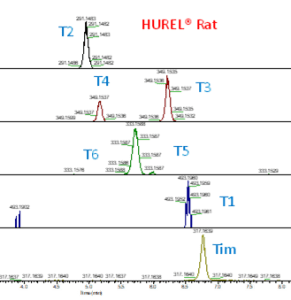
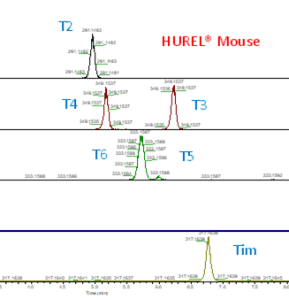
Figure 2: Chromatogram of Timolol and its metabolites in HUREL Rat and Mouse model
With these instruments and capabilities of HUREL microl iver assays, Visikol can provide an in vitro model akin to those of an in vivo model. Call or inquire today about a service we offer.
